networkx module¶
NetworkX¶
https://github.com/networkx/networkx
Aric A. Hagberg, Daniel A. Schult and Pieter J. Swart, “Exploring network structure, dynamics, and function using NetworkX”, in Proceedings of the 7th Python in Science Conference (SciPy2008), Gäel Varoquaux, Travis Vaught, and Jarrod Millman (Eds), (Pasadena, CA USA), pp. 11–15, Aug 2008 (pdf).
This content is originally downloaded from https://networkx.github.io/documentation/stable/tutorial.html and adapted to be shown as a presentation; moreover, we mix in additional resources such as examples (citing them and the original authors) in the last section.
Creating a graph¶
Create an empty graph with no nodes and no edges.
import networkx as nx
G = nx.Graph()
By definition, a Graph is a collection of nodes (vertices) along
with identified pairs of nodes (called edges, links, etc). In NetworkX,
nodes can be any hashable object e.g., a text string, an image, an XML
object, another Graph, a customized node object, etc.
An object is hashable if it has a hash value which never changes during its lifetime (it needs a
__hash__()method), and can be compared to other objects (it needs an__eq__()method). Hashable objects which compare equal must have the same hash value.
type(G)
networkx.classes.graph.Graph
Nodes¶
The graph G can be grown in several ways. NetworkX includes many
graph generator functions and facilities to read and write graphs in
many formats. To get started though we’ll look at simple manipulations.
You can add one node at a time,
G.add_node(1)
G.nodes
NodeView((1,))
add a list of nodes,
G.add_nodes_from([2, 3])
nx.draw(G)

help(G.add_nodes_from)
Help on method add_nodes_from in module networkx.classes.graph:
add_nodes_from(nodes_for_adding, **attr) method of networkx.classes.graph.Graph instance
Add multiple nodes.
Parameters
----------
nodes_for_adding : iterable container
A container of nodes (list, dict, set, etc.).
OR
A container of (node, attribute dict) tuples.
Node attributes are updated using the attribute dict.
attr : keyword arguments, optional (default= no attributes)
Update attributes for all nodes in nodes.
Node attributes specified in nodes as a tuple take
precedence over attributes specified via keyword arguments.
See Also
--------
add_node
Examples
--------
>>> G = nx.Graph() # or DiGraph, MultiGraph, MultiDiGraph, etc
>>> G.add_nodes_from("Hello")
>>> K3 = nx.Graph([(0, 1), (1, 2), (2, 0)])
>>> G.add_nodes_from(K3)
>>> sorted(G.nodes(), key=str)
[0, 1, 2, 'H', 'e', 'l', 'o']
Use keywords to update specific node attributes for every node.
>>> G.add_nodes_from([1, 2], size=10)
>>> G.add_nodes_from([3, 4], weight=0.4)
Use (node, attrdict) tuples to update attributes for specific nodes.
>>> G.add_nodes_from([(1, dict(size=11)), (2, {"color": "blue"})])
>>> G.nodes[1]["size"]
11
>>> H = nx.Graph()
>>> H.add_nodes_from(G.nodes(data=True))
>>> H.nodes[1]["size"]
11
or add any iterable container of nodes. You can also add nodes along with node attributes if your container yields 2-tuples (node, node_attribute_dict). Node attributes are discussed further below.
H = nx.path_graph(10)
nx.draw(H)
type(H)
networkx.classes.graph.Graph
G.add_nodes_from(H)
Note that G now contains the nodes of H as nodes of G. In
contrast, you could use the graph H as a node in G.
G.add_node(H)
The graph G now contains H as a node. This flexibility is very
powerful as it allows graphs of graphs, graphs of files, graphs of
functions and much more. It is worth thinking about how to structure
your application so that the nodes are useful entities. Of course you
can always use a unique identifier in G and have a separate
dictionary keyed by identifier to the node information if you prefer.
Edges¶
G can also be grown by adding one edge at a time,
G.add_edge(1, 2)
e = (2, 3)
G.add_edge(*e) # unpack edge tuple*
by adding a list of edges,
G.add_edges_from([(1, 2), (1, 3)])
or by adding any ebunch of edges. An ebunch is any iterable container
of edge-tuples. An edge-tuple can be a 2-tuple of nodes or a 3-tuple
with 2 nodes followed by an edge attribute dictionary, e.g.,
(2, 3, {'weight': 3.1415}). Edge attributes are discussed further
below
G.add_edges_from(H.edges)
G.edges
EdgeView([(1, 2), (1, 0), (2, 3), (3, 4), (4, 5), (5, 6), (6, 7), (7, 8), (8, 9)])
There are no complaints when adding existing nodes or edges. For example, after removing all nodes and edges,
G.clear()
len(G)
0
we add new nodes/edges and NetworkX quietly ignores any that are already present.
G.add_edges_from([(1, 2), (1, 3)])
G.add_node(1)
G.add_edge(1, 2)
G.add_node("spam") # adds node "spam"
G.add_nodes_from("spam") # adds 4 nodes: 's', 'p', 'a', 'm'
G.add_edge(3, 'm')
At this stage the graph G consists of 8 nodes and 3 edges, as can be
seen by:
G.number_of_nodes()
8
G.number_of_edges()
3
We can examine the nodes and edges. Four basic graph properties
facilitate reporting: G.nodes, G.edges, G.adj and
G.degree. These are set-like views of the nodes, edges, neighbors
(adjacencies), and degrees of nodes in a graph. They offer a continually
updated read-only view into the graph structure. They are also dict-like
in that you can look up node and edge data attributes via the views and
iterate with data attributes using methods .items(),
.data('span'). If you want a specific container type instead of a
view, you can specify one. Here we use lists, though sets, dicts, tuples
and other containers may be better in other contexts.
list(G.nodes)
[1, 2, 3, 'spam', 's', 'p', 'a', 'm']
list(G.edges)
[(1, 2), (1, 3), (3, 'm')]
list(G.adj[1]) # or list(G.neighbors(1))
[2, 3]
G.degree[1] # the number of edges incident to 1
2
One can specify to report the edges and degree from a subset of all nodes using an nbunch. An nbunch is any of: None (meaning all nodes), a node, or an iterable container of nodes that is not itself a node in the graph.
G.edges([2, 'm']), G.degree([2, 3])
(EdgeDataView([(2, 1), ('m', 3)]), DegreeView({2: 1, 3: 2}))
One can remove nodes and edges from the graph in a similar fashion to
adding. Use methods Graph.remove_node(),
Graph.remove_nodes_from(), Graph.remove_edge() and
Graph.remove_edges_from(), e.g.
G.remove_node(2)
G.remove_nodes_from("spam")
list(G.nodes)
[1, 3, 'spam']
G.remove_edge(1, 3)
When creating a graph structure by instantiating one of the graph classes you can specify data in several formats.
G.add_edge(1, 2)
H = nx.DiGraph(G) # create a DiGraph using the connections from G
list(H.edges())
[(1, 2), (2, 1)]
edgelist = [(0, 1), (1, 2), (2, 3)]
H = nx.Graph(edgelist)
What to use as nodes and edges¶
You might notice that nodes and edges are not specified as NetworkX
objects. This leaves you free to use meaningful items as nodes and
edges. The most common choices are numbers or strings, but a node can be
any hashable object (except None), and an edge can be associated
with any object x using G.add_edge(n1, n2, object=x).
As an example, n1 and n2 could be protein objects from the RCSB
Protein Data Bank, and x could refer to an XML record of
publications detailing experimental observations of their interaction.
We have found this power quite useful, but its abuse can lead to
unexpected surprises unless one is familiar with Python. If in doubt,
consider using convert_node_labels_to_integers() to obtain a more
traditional graph with integer labels.
Accessing edges and neighbors¶
In addition to the views Graph.edges(), and Graph.adj(), access
to edges and neighbors is possible using subscript notation.
G[1] # same as G.adj[1]
AtlasView({2: {}})
G[1][2], G.edges[1, 2]
({}, {})
You can get/set the attributes of an edge using subscript notation if the edge already exists.
G.add_edge(1, 3)
G[1][3]['color'] = "blue"
G.edges[1, 2]['color'] = "red"
Fast examination of all (node, adjacency) pairs is achieved using
G.adjacency(), or G.adj.items(). Note that for undirected
graphs, adjacency iteration sees each edge twice.
FG = nx.Graph()
FG.add_weighted_edges_from(
[(1, 2, 0.125), (1, 3, 0.75), (2, 4, 1.2), (3, 4, 0.375)])
for n, nbrs in FG.adj.items():
for nbr, eattr in nbrs.items():
wt = eattr['weight']
if wt < 0.5: print('(%d, %d, %.3f)' % (n, nbr, wt))
(1, 2, 0.125)
(2, 1, 0.125)
(3, 4, 0.375)
(4, 3, 0.375)
Convenient access to all edges is achieved with the edges property.
for (u, v, wt) in FG.edges.data('weight'):
if wt < 0.5: print('(%d, %d, %.3f)' % (u, v, wt))
(1, 2, 0.125)
(3, 4, 0.375)
Adding attributes to graphs, nodes, and edges¶
Attributes such as weights, labels, colors, or whatever Python object you like, can be attached to graphs, nodes, or edges.
Each graph, node, and edge can hold key/value attribute pairs in an
associated attribute dictionary (the keys must be hashable). By default
these are empty, but attributes can be added or changed using
add_edge, add_node or direct manipulation of the attribute
dictionaries named G.graph, G.nodes, and G.edges for a graph
G.
Graph attributes¶
Assign graph attributes when creating a new graph
G = nx.Graph(day="Friday")
G.graph
{'day': 'Friday'}
type(_)
dict
Or you can modify attributes later
G.graph['day'] = "Monday"
G.graph
{'day': 'Monday'}
Add node attributes using add_node(), add_nodes_from(), or
G.nodes
G.add_node(1, time='5pm')
G.add_nodes_from([3], time='2pm')
G.nodes[1]
{'time': '5pm'}
G.nodes[1]['room'] = 714
G.nodes.data()
NodeDataView({1: {'time': '5pm', 'room': 714}, 3: {'time': '2pm'}})
Note that adding a node to G.nodes does not add it to the graph, use
G.add_node() to add new nodes. Similarly for edges.
Add/change edge attributes using add_edge(), add_edges_from(),
or subscript notation.
G.add_edge(1, 2, weight=4.7 )
G.add_edges_from([(3, 4), (4, 5)], color='red')
G.add_edges_from([(1, 2, {'color': 'blue'}), (2, 3, {'weight': 8})])
G[1][2]['weight'] = 4.7
G.edges[3, 4]['weight'] = 4.2
The special attribute weight should be numeric as it is used by
algorithms requiring weighted edges.
Directed graphs
The DiGraph class provides additional properties specific to
directed edges, e.g., DiGraph.out_edges(), DiGraph.in_degree(),
DiGraph.predecessors(), DiGraph.successors() etc. To allow
algorithms to work with both classes easily, the directed versions of
neighbors() is equivalent to successors() while degree
reports the sum of in_degree and out_degree even though that may
feel inconsistent at times.
DG = nx.DiGraph()
DG.add_weighted_edges_from([(1, 2, 0.5), (3, 1, 0.75)])
DG.out_degree(1, weight='weight'), DG.degree(1, weight='weight')
(0.5, 1.25)
list(DG.successors(1)), list(DG.neighbors(1))
([2], [2])
Some algorithms work only for directed graphs and others are not well
defined for directed graphs. Indeed the tendency to lump directed and
undirected graphs together is dangerous. If you want to treat a directed
graph as undirected for some measurement you should probably convert it
using Graph.to_undirected() or with
H = nx.Graph(G) # convert G to undirected graph
NetworkX provides classes for graphs which allow multiple edges between
any pair of nodes. The MultiGraph and MultiDiGraph classes allow
you to add the same edge twice, possibly with different edge data. This
can be powerful for some applications, but many algorithms are not well
defined on such graphs. Where results are well defined, e.g.,
MultiGraph.degree() we provide the function. Otherwise you should
convert to a standard graph in a way that makes the measurement well
defined.
MG = nx.MultiGraph()
MG.add_weighted_edges_from([(1, 2, 0.5), (1, 2, 0.75), (2, 3, 0.5)])
dict(MG.degree(weight='weight'))
{1: 1.25, 2: 1.75, 3: 0.5}
GG = nx.Graph()
for n, nbrs in MG.adjacency():
for nbr, edict in nbrs.items():
minvalue = min([d['weight'] for d in edict.values()])
GG.add_edge(n, nbr, weight = minvalue)
nx.shortest_path(GG, 1, 3)
[1, 2, 3]
In addition to constructing graphs node-by-node or edge-by-edge, they can also be generated by
Applying classic graph operations, such as:
subgraph(G, nbunch) - induced subgraph view of G on nodes in nbunch union(G1,G2) - graph union disjoint_union(G1,G2) - graph union assuming all nodes are different cartesian_product(G1,G2) - return Cartesian product graph compose(G1,G2) - combine graphs identifying nodes common to both complement(G) - graph complement create_empty_copy(G) - return an empty copy of the same graph class to_undirected(G) - return an undirected representation of G to_directed(G) - return a directed representation of G
Using a call to one of the classic small graphs, e.g.,
petersen = nx.petersen_graph()
nx.draw(petersen)
tutte = nx.tutte_graph()
nx.draw(tutte)
maze = nx.sedgewick_maze_graph()
nx.draw(maze)
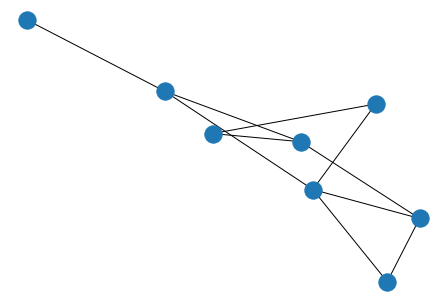
tet = nx.tetrahedral_graph()
nx.draw(tet)
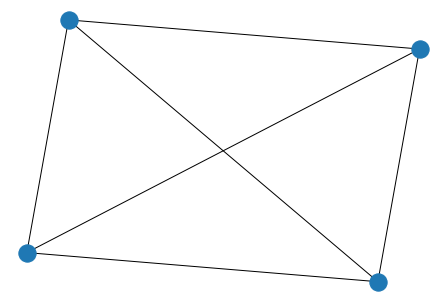
K_5 = nx.complete_graph(5)
nx.draw(K_5)
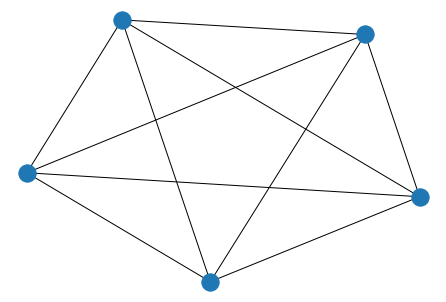
K_3_5 = nx.complete_bipartite_graph(3, 5)
nx.draw(K_3_5)
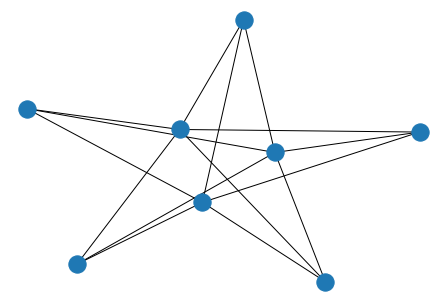
barbell = nx.barbell_graph(10, 10)
nx.draw(barbell)
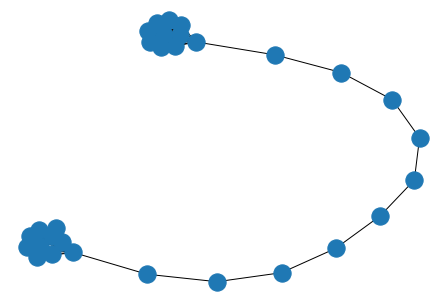
lollipop = nx.lollipop_graph(10, 20)
nx.draw(lollipop)
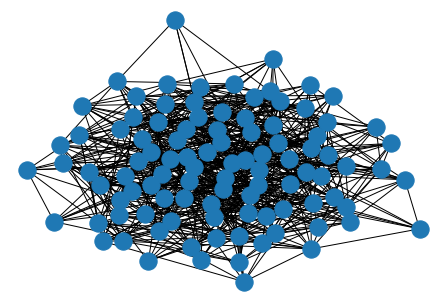
er = nx.erdos_renyi_graph(100, 0.15)
nx.draw(er)
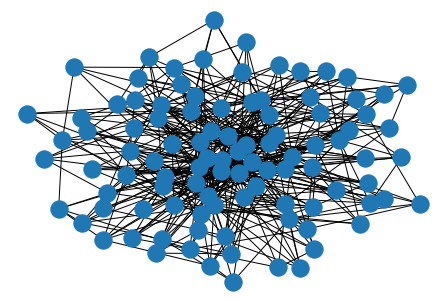
ws = nx.watts_strogatz_graph(30, 3, 0.1)
nx.draw(ws)

ba = nx.barabasi_albert_graph(100, 5)
nx.draw(ba)
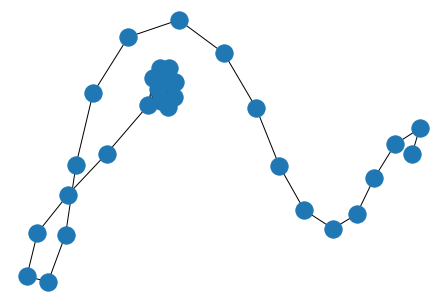
red = nx.random_lobster(100, 0.9, 0.9)
nx.draw(red)
nx.write_gml(red, "path.to.file")
mygraph = nx.read_gml("path.to.file")
nx.draw(mygraph)
The structure of G can be analyzed using various graph-theoretic
functions such as:
G = nx.Graph()
G.add_edges_from([(1, 2), (1, 3)])
G.add_node("spam") # adds node "spam"
list(nx.connected_components(G))
[{1, 2, 3}, {'spam'}]
sorted(d for n, d in G.degree())
[0, 1, 1, 2]
nx.clustering(G)
{1: 0, 2: 0, 3: 0, 'spam': 0}
Some functions with large output iterate over (node, value) 2-tuples. These are easily stored in a dict structure if you desire.
sp = dict(nx.all_pairs_shortest_path(G))
sp[3]
{3: [3], 1: [3, 1], 2: [3, 1, 2]}
See Algorithms for details on graph algorithms supported.
NetworkX is not primarily a graph drawing package but basic drawing with
Matplotlib as well as an interface to use the open source Graphviz
software package are included. These are part of the
networkx.drawing module and will be imported if possible.
First import Matplotlib’s plot interface (pylab works too)
import matplotlib.pyplot as plt
You may find it useful to interactively test code using
ipython -pylab, which combines the power of ipython and matplotlib
and provides a convenient interactive mode.
To test if the import of networkx.drawing was successful draw G
using one of
G = nx.petersen_graph()
plt.subplot(121)
nx.draw(G, with_labels=True, font_weight='bold')
plt.subplot(122)
nx.draw_shell(G, nlist=[range(5, 10), range(5)],
with_labels=True, font_weight='bold')
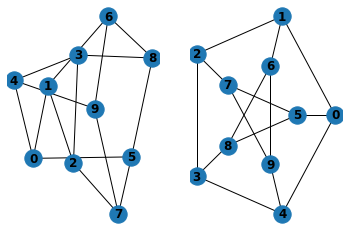
when drawing to an interactive display. Note that you may need to issue a Matplotlib
plt.show()
command if you are not using matplotlib in interactive mode (see Matplotlib FAQ ).
options = {
'node_color': 'black',
'node_size': 100,
'width': 3,
}
plt.subplot(221)
nx.draw_random(G, **options)
plt.subplot(222)
nx.draw_circular(G, **options)
plt.subplot(223)
nx.draw_spectral(G, **options)
plt.subplot(224)
nx.draw_shell(G, nlist=[range(5,10), range(5)], **options)
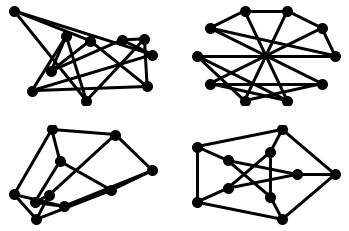
You can find additional options via draw_networkx() and layouts via
layout. You can use multiple shells with draw_shell().
G = nx.dodecahedral_graph()
shells = [[2, 3, 4, 5, 6],
[8, 1, 0, 19, 18, 17, 16, 15, 14, 7],
[9, 10, 11, 12, 13]]
nx.draw_shell(G, nlist=shells, **options)
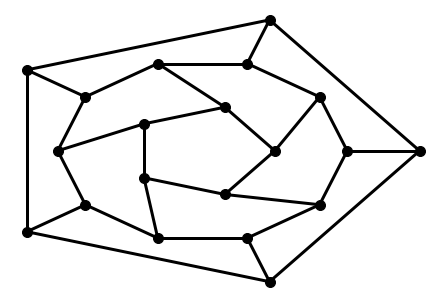
To save drawings to a file, use, for example
nx.draw(G)
plt.savefig("path.png")
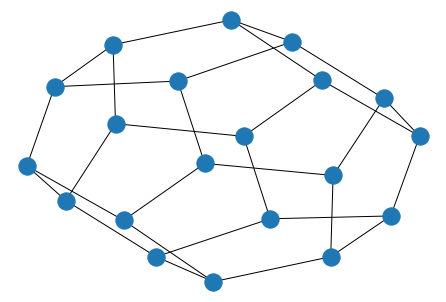
writes to the file path.png in the local directory.
If Graphviz and PyGraphviz or pydot, are available on your system, you
can also use nx_agraph.graphviz_layout(G) or
nx_pydot.graphviz_layout(G) to get the node positions, or write the
graph in dot format for further processing.
from networkx.drawing.nx_pydot import write_dot
pos = nx.nx_agraph.graphviz_layout(G)
nx.draw(G, pos=pos)
write_dot(G, 'file.dot')
See Drawing for additional details.
A complete gallery of examples can be found at https://networkx.github.io/documentation/stable/auto_examples/index.html
# Author: Aric Hagberg (hagberg@lanl.gov)
G = nx.grid_2d_graph(5, 5) # 5x5 grid
# print the adjacency list
#for line in nx.generate_adjlist(G):
# print(line)
# write edgelist to grid.edgelist
nx.write_edgelist(G, path="grid.edgelist", delimiter=":")
# read edgelist from grid.edgelist
H = nx.read_edgelist(path="grid.edgelist", delimiter=":")
nx.draw(H)
plt.show()
G = nx.lollipop_graph(4, 6)
pathlengths = []
print("source vertex {target:length, }")
for v in G.nodes():
spl = dict(nx.single_source_shortest_path_length(G, v))
print('{} {} '.format(v, spl))
for p in spl:
pathlengths.append(spl[p])
print('')
print("average shortest path length %s" %
(sum(pathlengths) / len(pathlengths)))
source vertex {target:length, }
0 {0: 0, 1: 1, 2: 1, 3: 1, 4: 2, 5: 3, 6: 4, 7: 5, 8: 6, 9: 7}
1 {1: 0, 0: 1, 2: 1, 3: 1, 4: 2, 5: 3, 6: 4, 7: 5, 8: 6, 9: 7}
2 {2: 0, 0: 1, 1: 1, 3: 1, 4: 2, 5: 3, 6: 4, 7: 5, 8: 6, 9: 7}
3 {3: 0, 0: 1, 1: 1, 2: 1, 4: 1, 5: 2, 6: 3, 7: 4, 8: 5, 9: 6}
4 {4: 0, 5: 1, 3: 1, 6: 2, 0: 2, 1: 2, 2: 2, 7: 3, 8: 4, 9: 5}
5 {5: 0, 4: 1, 6: 1, 3: 2, 7: 2, 0: 3, 1: 3, 2: 3, 8: 3, 9: 4}
6 {6: 0, 5: 1, 7: 1, 4: 2, 8: 2, 3: 3, 9: 3, 0: 4, 1: 4, 2: 4}
7 {7: 0, 6: 1, 8: 1, 5: 2, 9: 2, 4: 3, 3: 4, 0: 5, 1: 5, 2: 5}
8 {8: 0, 7: 1, 9: 1, 6: 2, 5: 3, 4: 4, 3: 5, 0: 6, 1: 6, 2: 6}
9 {9: 0, 8: 1, 7: 2, 6: 3, 5: 4, 4: 5, 3: 6, 0: 7, 1: 7, 2: 7}
average shortest path length 2.86
# histogram of path lengths
dist = {}
for p in pathlengths:
if p in dist:
dist[p] += 1
else:
dist[p] = 1
print('')
print("length #paths")
verts = dist.keys()
for d in sorted(verts):
print('%s %d' % (d, dist[d]))
length #paths
0 10
1 24
2 16
3 14
4 12
5 10
6 8
7 6
print("radius: %d" % nx.radius(G))
print("diameter: %d" % nx.diameter(G))
print("eccentricity: %s" % nx.eccentricity(G))
print("center: %s" % nx.center(G))
print("periphery: %s" % nx.periphery(G))
print("density: %s" % nx.density(G))
radius: 4
diameter: 7
eccentricity: {0: 7, 1: 7, 2: 7, 3: 6, 4: 5, 5: 4, 6: 4, 7: 5, 8: 6, 9: 7}
center: [5, 6]
periphery: [0, 1, 2, 9]
density: 0.26666666666666666
nx.draw(G, with_labels=True)
plt.show()
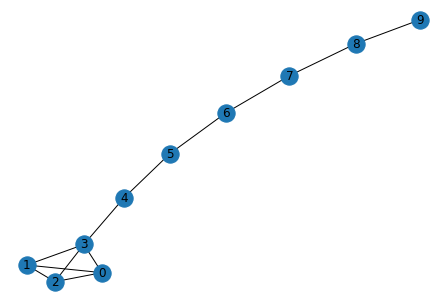
# Author: Aric Hagberg (hagberg@lanl.gov)
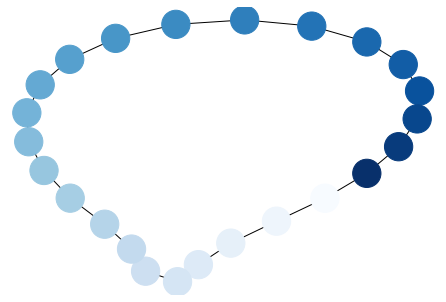
G = nx.cycle_graph(24)
pos = nx.spring_layout(G, iterations=200)
nx.draw(G, pos, node_color=range(24),
node_size=800, cmap=plt.cm.Blues)
plt.show()
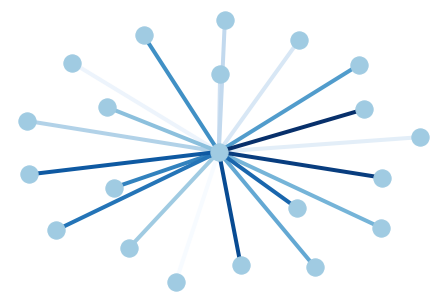
# Author: Aric Hagberg (hagberg@lanl.gov)
G = nx.star_graph(20)
pos = nx.spring_layout(G)
colors = range(20)
nx.draw(G, pos, node_color='#A0CBE2', edge_color=colors,
width=4, edge_cmap=plt.cm.Blues, with_labels=False)
plt.show()
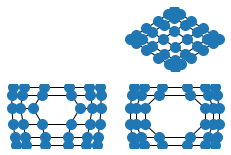
options = {
'node_color': 'C0',
'node_size': 100,
}
G = nx.grid_2d_graph(6, 6)
plt.subplot(332)
nx.draw_spectral(G, **options)
G.remove_edge((2, 2), (2, 3))
plt.subplot(334)
nx.draw_spectral(G, **options)
G.remove_edge((3, 2), (3, 3))
plt.subplot(335)
nx.draw_spectral(G, **options)
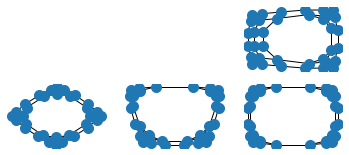
G.remove_edge((2, 2), (3, 2))
plt.subplot(336)
nx.draw_spectral(G, **options)
G.remove_edge((2, 3), (3, 3))
plt.subplot(337)
nx.draw_spectral(G, **options)
G.remove_edge((1, 2), (1, 3))
plt.subplot(338)
nx.draw_spectral(G, **options)
G.remove_edge((4, 2), (4, 3))
plt.subplot(339)
nx.draw_spectral(G, **options)
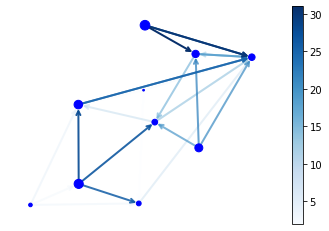
# Author: Rodrigo Dorantes-Gilardi (rodgdor@gmail.com)
import matplotlib as mpl
import matplotlib.pyplot as plt
import networkx as nx
G = nx.generators.directed.random_k_out_graph(10, 3, 0.5)
pos = nx.layout.spring_layout(G)
node_sizes = [3 + 10 * i for i in range(len(G))]
M = G.number_of_edges()
edge_colors = range(2, M + 2)
edge_alphas = [(5 + i) / (M + 4) for i in range(M)]
nodes = nx.draw_networkx_nodes(G, pos, node_size=node_sizes, node_color='blue')
edges = nx.draw_networkx_edges(G, pos, node_size=node_sizes, arrowstyle='->',
arrowsize=10, edge_color=edge_colors,
edge_cmap=plt.cm.Blues, width=2)
# set alpha value for each edge
for i in range(M):
edges[i].set_alpha(edge_alphas[i])
pc = mpl.collections.PatchCollection(edges, cmap=plt.cm.Blues)
pc.set_array(edge_colors)
plt.colorbar(pc)
ax = plt.gca()
ax.set_axis_off()
plt.show()
# Authors: Brendt Wohlberg, Aric Hagberg (hagberg@lanl.gov)
import gzip
import re
import sys
import matplotlib.pyplot as plt
from networkx import nx
def roget_graph():
""" Return the thesaurus graph from the roget.dat example in
the Stanford Graph Base.
"""
# open file roget_dat.txt.gz (or roget_dat.txt)
fh = gzip.open('roget_dat.txt.gz', 'r')
G = nx.DiGraph()
for line in fh.readlines():
line = line.decode()
if line.startswith("*"): # skip comments
continue
if line.startswith(" "): # this is a continuation line, append
line = oldline + line
if line.endswith("\\\n"): # continuation line, buffer, goto next
oldline = line.strip("\\\n")
continue
(headname, tails) = line.split(":")
# head
numfind = re.compile("^\d+") # re to find the number of this word
head = numfind.findall(headname)[0] # get the number
G.add_node(head)
for tail in tails.split():
if head == tail:
print("skipping self loop", head, tail, file=sys.stderr)
G.add_edge(head, tail)
return G
G = roget_graph()
print("Loaded roget_dat.txt containing 1022 categories.")
print("digraph has %d nodes with %d edges"
% (nx.number_of_nodes(G), nx.number_of_edges(G)))
UG = G.to_undirected()
print(nx.number_connected_components(UG), "connected components")
options = {
'node_color': 'black',
'node_size': 1,
'line_color': 'grey',
'linewidths': 0,
'width': 0.1,
}
nx.draw_circular(UG, **options)
plt.show()
skipping self loop 400 400
Loaded roget_dat.txt containing 1022 categories.
digraph has 1022 nodes with 5075 edges
21 connected components